
Read this post on the Asimov website. Read Part I.
Strings of DNA
Every cell in the human body has the same genome. There are slight variations between each one — so-called somatic mutations — but the basic gist holds true.
This raises a difficult question: If every genome in every cell is (basically) the same, what accounts for the remarkable diversity of form and function in a single person? A neuron fires action potentials up to 100 times per second; the signals ripple through the brain at speeds exceeding 500 miles per hour. Osteoclast cells break down bones and can contain up to 100 nuclei, instead of just one. Skin cells in the feet make toenails, but skin on the elbow doesn’t.
How does a cell know what it is, or what it does?
The answer is genome regulation. Individual cell types form during development and then become rooted — fixed — in their identities. Each cell type coils up its genome in a distinct way. Large segments of genes are ‘shut down’ in neurons, but those same genes remain open and active in toenail-making cells. Every genome also has epigenetic differences; chemical tags (like acetyl or methyl groups) tune gene expression.
In short, two cells with identical DNA will package and read the genetic code in different ways.
It’s not yet possible to look at an arbitrary genome region and say, “this will be active in the retina,” or “this will be active in the heart.” But we are genetic designers, and we want to reliably engineer biology. By designing tools that can analyze a DNA sequence and predict its expression across the body, it becomes possible to also do the reverse: design custom DNA that is active only in specific tissues. Such a tool would be transformative for gene therapies.
Our previous blog post explained how viral vectors are packaged with DNA and injected into the human body to treat disease. Gene therapies are a molecular marvel, but they also have problems. Viral vectors often transduce cells that we don’t want them to. A gene therapy for the brain often ends up in the liver. These off-target effects can cause side effects and are sometimes lethal. By packaging gene therapies with DNA sequences that only ‘switch on’ in specific tissues, though, we can improve their safety and efficacy.
In this blog, we explain how we are using transformer-based AI models to study gene regulation and then design synthetic promoters that accomplish this aim.
AI-Guided Design
There are about 20,000 protein-coding genes in the human genome. Deciphering how all of them switch ‘on’ and ‘off’ has challenged scientists for the last six decades. So let’s focus, instead, on just one gene.
In human cells, regulatory sequences surround the coding sequence that encodes a protein. Upstream of the gene, a promoter acts as an ignition switch for transcription. Downstream, a terminator signals the end of transcription. Enhancer sequences boost a gene’s expression and can appear anywhere, upstream or downstream, even millions of bases away.
This model of a gene is simplistic. It makes it sound as if we’ve got everything figured out! But even in cases where we know the identities of all the proteins that interact with a gene, and have mapped the sequence and position of every element — promoters, terminators, enhancers, and more — we still don’t fully understand how all the molecules come together and actually dance to control a gene’s expression.
AI tools can help. They’re already used to study complex phenomena, such as computer vision or natural language. But perhaps they can also predict how genes are regulated, and thus make it possible to design new promoters that behave in desired ways.
Consider transformers, the same type of neural network underlying ChatGPT. Originally developed to translate languages in 2017, transformers quickly surpassed recurrent neural networks in their ability to both translate language and infer the semantic connections between words.
Whereas a recurrent neural network ‘reads’ and ‘processes’ one word at a time — and quickly forgets words as it moves along a paragraph — a transformer can be parallelized across many words while learning and tracking the semantic connections between each one.
Consider two sentences: “Server, can I have the check?” and “Looks like I just crashed the server.” The word server appears in both sentences, but it has a different meaning in each case. The intended meaning of the word can be deciphered by looking at adjacent words, such as check in the first sentence and crashed in the second. Transformers pick up on these nuances.
We are fortunate, in a way, that biological sequences are reminiscent of written languages. DNA and proteins are represented as strings of letters, and these letters have regulatory and functional meanings. The letters TATAAA in a DNA sequence has a physical meaning: It’s a binding site for transcription factors that activate or repress a gene. The similarities between words and DNA mean that transformers can be brought to bear on biology.
AlphaFold used a transformer to predict 200 million protein structures from strings of amino acids. Other groups have used transformers to predict gene expression from DNA sequences or to predict which proteins will get ‘tagged’ with sugars in a cell.
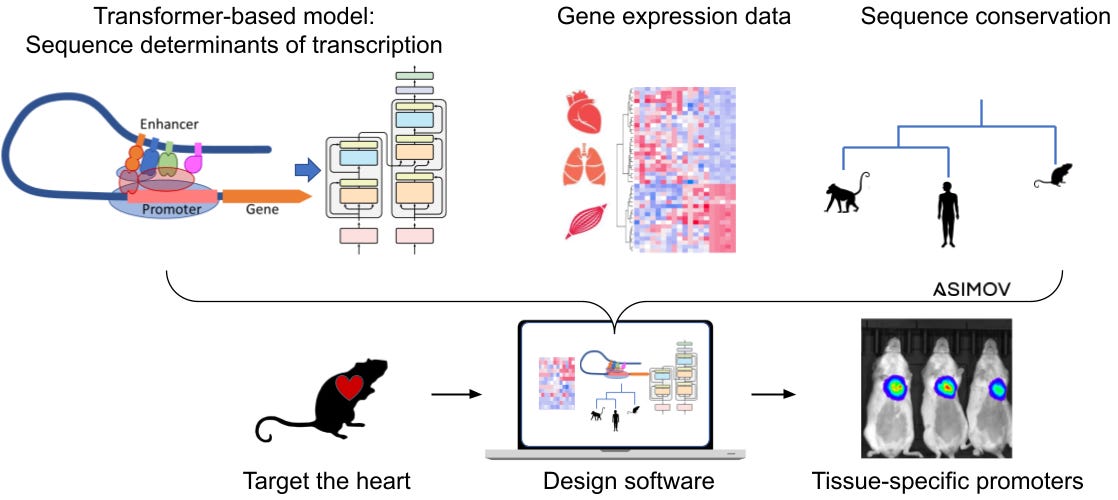
At Asimov, we recently developed a computational tool to design synthetic promoters that are only active in specific tissues. The computational tool merges a transformer-based model, tissue RNA-sequencing datasets (a measure of gene expression in different parts of the body), and evolutionary data on sequence conservation (i.e. gene sequences that appear in both mice and humans).
The generated sequences successfully confined gene expression to our target tissues. And although the synthetic promoters do not naturally exist in nature, the evolutionary data used in their design means that they are effectively "humanized." We expect that these promoters will translate from mice into humans without a loss of function.
The Experiment
Using our transformer-based design tool, our engineering team modeled and synthesized 17 synthetic promoters to target the heart, brain, liver, or muscles. Each promoter was fused to a luciferase coding sequence. The luciferase protein emits light, and it’s the same molecule that fireflies use to flash and communicate.
Luciferase will emit light in the presence of luciferin, ATP, magnesium, and oxygen. When we add luciferin to an animal’s drinking water, their organs literally glow if the luciferase payload is expressed within. We chose to use luciferase in these experiments because it has a higher sensitivity than fluorescent reporters. We are able to measure luciferase even if it is only weakly expressed.
Each DNA sequence — a promoter, luciferase coding gene, terminator, and other bits and pieces — was packaged into a recombinant AAV9 capsid to simulate their delivery as a gene therapy. An AAV is made from two parts: a 20-sided icosahedral protein shell, called a capsid, and up to 4,700 bases of single-stranded DNA packed inside.
Three mice, all with an identical genetic background, were intravenously injected with the engineered AAVs. We used two well-studied promoters as controls: CMV, which is understood to express in all tissues, and MHCK7, which expresses in both heart and muscle.
Every animal was given the same dosage of viral vector: 6 x 1013 viral genomes per kilogram of body mass. (A typical mouse weighs less than one ounce, so the animals were each injected with a few billion viral vectors.)
And then, we waited.
After two weeks, we imaged each animal and directly measured the bioluminescence of each tissue using a camera that counts photons. In the fourth week, we repeated the imaging and collected tissues for DNA quantification and RT-qPCR, a technique that counts the number of mRNA transcripts for a gene. We divided the total number of RNAs for luciferase by the number for GAPDH, a protein that has consistent gene expression throughout the day. We repeated this for every mouse and every promoter.
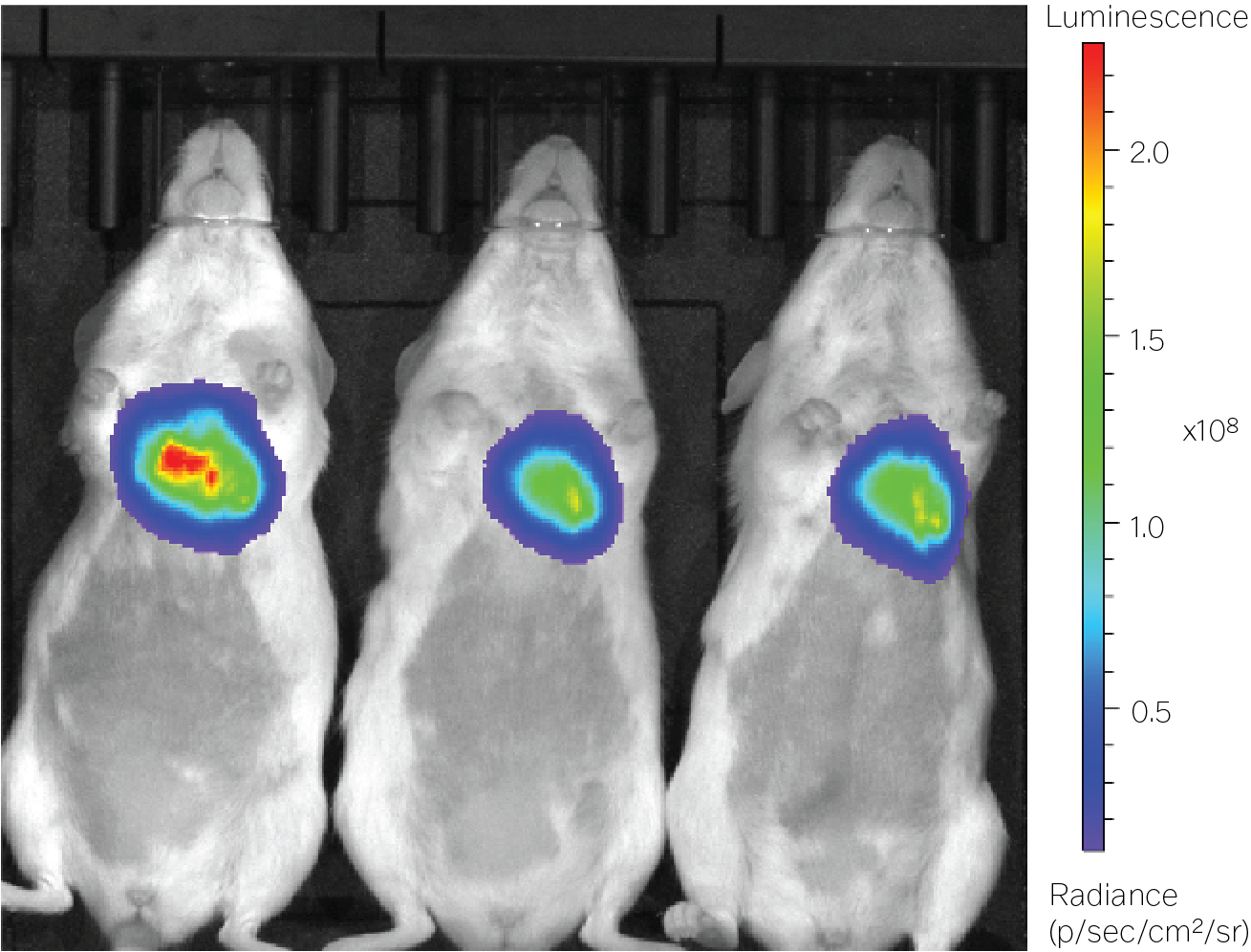
The synthetic promoter that we designed for the heart worked as expected. The luciferase imaging data and RNA quantification results both indicated that the promoter expressed luciferase at levels 120-times higher in the heart compared with the liver (see the Heart panel in the figure.)

The packaging limit of a recombinant AAV is quite small — just 4,700 bases of DNA — so we also wanted to see whether it’d be possible to truncate, or shorten, our synthetic promoter while maintaining its specificity for the heart. And it was. A shortened version of the heart-specific promoter had 29-times higher expression in the heart versus liver (see the Heart / Truncated panel in the figure), even when we cut the promoter to 40 percent of its original length.
Doubling the gene therapy’s dose also did not substantially increase the promoter’s expression in off-target tissues. It resulted, instead, in 208-times higher expression in the heart versus liver (see the High dose panel). We’ve already taken these observations and fed them back into our model to improve predictions for future design cycles.
Results for other promoters were equally intriguing.
A promoter designed to express in the muscles, for example, did not work at all. There was no bioluminescence and very little RNA expression (see the Muscle panel). But when we chopped off a small part of the promoter’s sequence, it became 37-times more active in the intestines compared to the liver (see the Muscle / Truncated panel.) We don’t fully understand why this happened, but the intestine contains another type of muscle: smooth muscle. Could this result be due to our synthetic promoter getting its wires crossed between these two muscle types? We’re not sure.
The transformer model did remarkably well overall. Computational tools don’t need to be perfect every time. They just need to be good enough to increase the probability of getting a winner within the confines of an experimental budget (a few dozen mice, a 96-well plate, and so on).
A promoter sequence that is 100 bases in length has 4100 possible permutations. This number far exceeds the number of atoms in the universe. Our transformer-based design tool shrinks the space of combinatorial possibilities. It finds winners faster than random screens or high-throughput experiments. And, we think, it will help us create better gene therapies.
If you’re excited about the intersection of AI and biology, come join us. Or, subscribe for updates.
Contributors: Alina Ferdman, Ben Gordon, Will Johnson, Alec Nielsen, Raja Srinivas, Stephen von Stetina & and Dinghai Zheng. Text by Niko McCarty.


I don't think this article is well-written because you didn't provide clear instructions on how to use the transformer model to design synthetic promoters, which should be the most valuable part of the article. Instead, the lengthy descriptions of the experimental process seem to be just routine validation without any novelty.