This is Part 1 in a series on the tools Asimov is building to engineer and model mammalian cells. It’s meant to be a broad introduction to our ideas. Consider reading this article on our (new) website.
In 1948, a rodent breeder in New York named Victor Schwentker sent a message to a colleague in China: Could you get some hamsters for me?
Schwentker had heard tales of a special hamster, native to northern China and Mongolia, with short gestation periods and natural resistance to human viruses — traits that would make them ideal for scientific research. Chinese scientists had used these hamsters to study pathogens since the 1910s, but Schwentker feared that the ongoing Chinese civil war would soon make them impossible to retrieve.
In the waning months of 1948, Schwentker sent a letter to Robert Briggs Watson, a Rockefeller Foundation field staff member, and asked him to bring (or, erm, smuggle) some hamsters back to New York.
Watson collected twenty hamsters – ten males and ten females, packed in a wooden crate – with help from a Chinese physician. He smuggled them out of the country via a Pan-Am flight from Shanghai, just before Communists took control of the country. (An excellent account of this story can be read online.)
Back in New York, Schwentker bred the hamsters and distributed them to other researchers, one of whom was a geneticist named Theodore Puck. In 1957, Puck took a small piece from a hamster’s ovary, plated the cells onto a dish, and passaged them. He isolated a clone that could divide again and again; an “immortalized” Chinese Hamster Ovary (CHO) cell with a genetic mutation that rendered them immune to normal senescence. A decade later, Puck also isolated a second clone that was unable to synthesize proline, an amino acid. Those cells could only be grown in media with added proline, thus paving the way for selection methods.
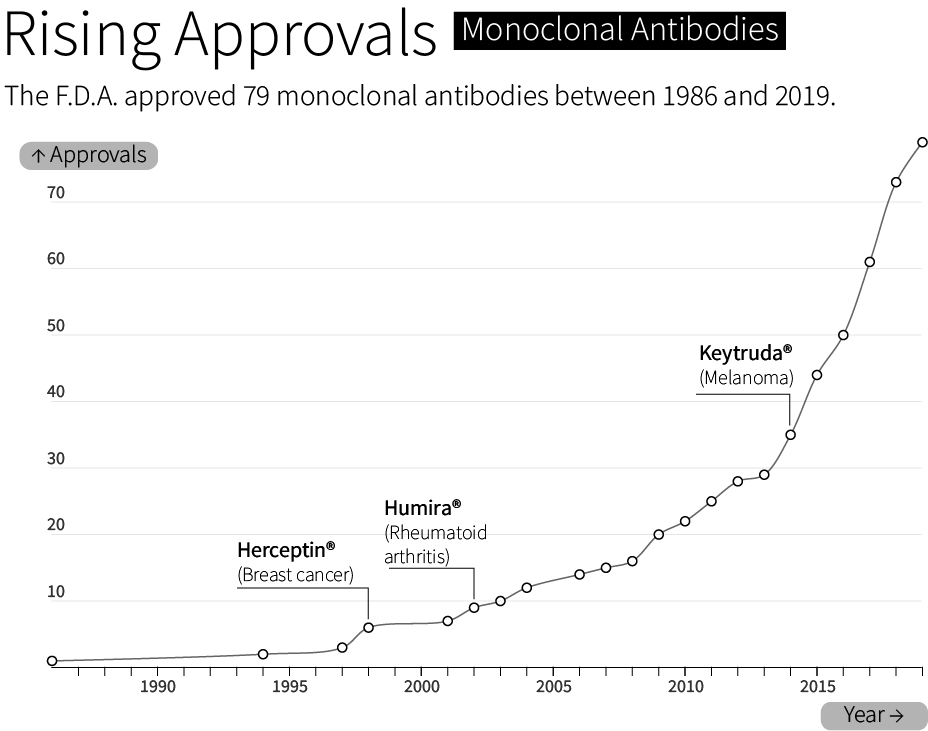
By 1986, the F.D.A. had approved the first “recombinant biotherapeutic protein” made by CHO cells. Today, CHO cells make roughly 70 percent of all F.D.A. approved biologics — molecules made by living cells — sold on the market. This includes bestsellers like Humira®, whose sales exceeded $21 billion in 2022, and Keytruda®, a cancer therapy with $20.9 billion sold in the same year.
It is remarkable, in a way, that a cell derived from a hamster, smuggled out of China halfway through the 20th century, has dominated biologics manufacturing for the last forty-odd years. But there’s a simple reason for its success: CHO cells are really good at making medicines.
They make lots of protein, routinely reaching production titers of multiple grams per liter. They also divide slightly faster than most human cells (18 to 24 hours) and grow at high densities, which means more cell ‘factories’ can be packed into the same volume. And CHO cells modify proteins in ways that microbial cells cannot; they tag proteins with sugars, or create disulfide bonds in proteins, much like human cells do.
CHO cells are also somewhat — but not always! — resistant to human viruses. There have been at least two cases where a rogue virus shut down a biologics factory. Genzyme facilities in Belgium and Allston, Massachusetts were infected with Vesivirus 2117, an RNA-packed, icosahedral virus, that cost the company “$100–300 million in lost sales,” according to reporting in Nature Biotechnology in 2009. (Other cell lines, such as NS0 and SP2/0, have also been used to make medicines, but they can add alpha-gal residues to proteins that cause severe immune reactions in people.)
Although CHO cells make many types of molecules, they are especially good at making monoclonal antibodies; ‘Y’ shaped proteins made from interwoven subunits, called heavy chains and light chains. Antibodies bind to molecules and proteins in the body with exceedingly high specificity. They are part of the body’s natural immune system, but are also used to treat everything from inflammation (Humira®) to COVID-19 (Actemra®). About 40% of all drugs are biologics, and antibodies alone will account for an estimated $300 billion in sales by 2025.

Coaxing CHO cells to make antibodies has followed basically the same recipe for the last 20 years. Each protein chain is first encoded as a DNA sequence and then placed downstream from a promoter that ‘drives’ its expression (historically, a promoter from human cytomegalovirus, called CMV). The DNA ’payload,’ encoding the antibody subunits, integrates into the genome. The cells begin to make antibodies.
Now here’s the problem. In the last two decades, individual pharma companies have relied on static methods — same genetic parts, same plasmid architecture, same host cells — to make antibodies with CHO cells. We now know, though, that the ratio of heavy and light chains, the number of DNA copies integrated into the genome, and many other factors in a dizzying constellation of possible “design space” can influence how much antibody a cell makes. The same factors can also determine how “good” the antibodies are: What fraction fold properly? Do the antibodies ‘clump’ up? Do they carry the correct sugar tags?
This is really important to understand, because the normal way to make antibodies today is with a “one-size-fits-all” approach. And that doesn’t work for every antibody.
If you were to take two different antibody sequences, place them in otherwise identical plasmids – with the same promoters and terminators – and drop them into CHO cells with identical genomes, those two antibodies might be produced at vastly different titers. In other words, the same plasmid doesn’t work for every antibody.
Even the codons within a DNA sequence (CUC, UUG and CUG all encode leucine, for example) can influence how many antibodies a cell makes. And these effects can be extreme; we’ve personally seen 3x boosts in antibody titers from codon optimization alone.
We suspect that it’s possible to boost both the quality and amount of antibodies that a cell makes, from a few grams per liter to more than 10 grams per liter, for even difficult-to-make molecules. And we aim to do that by building synthetic biology and computational models that can more efficiently explore the “genetic design space” of living cells. We call this technology stack for genetic design — which works across multiple cell types and applications, including CHO — our “CHO Edge” system.
CHO Edge includes different host cell lines and a library of thousands of characterized genetic elements — promoters, terminators, and many other “parts.” It also includes computational tools and a hyperactive transposase that tunably integrates between 20 and 60 copies of the plasmid into the CHO genome. (A transposase is an enzyme that cuts out a DNA sequence and then “pastes” it at sites across the genome.)
We’ve also made a custom codon optimizer and signal peptide predictor. Signal peptides are short sequences that cause a protein to be exported, or secreted, from the cell. Our tool predicts how well a given signal peptide will work with a given antibody, and we’ll explain how it works in an upcoming blog.
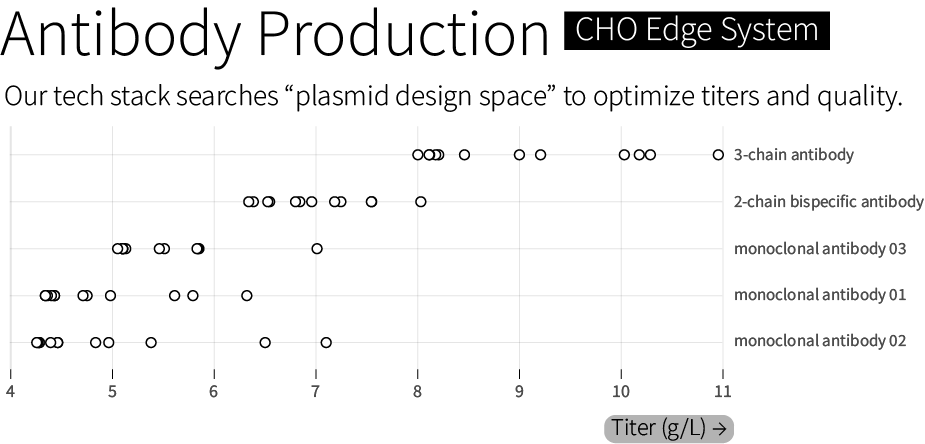
All of these tools are used, together, to search through “genetic design space” and optimize therapeutics manufacturing. We routinely achieve titers between 6 and 10 grams of antibody per liter. In one case, we achieved 11 grams per liter for a 3-chain bispecific antibody — a more complex protein architecture than standard 2-chain monoclonal antibodies — with 85 percent heterodimerization, a measure of a molecule’s quality. We guarantee a minimal titer of 4 grams per liter, or our work is free.
Boosted titers are useful not just because they cut down on manufacturing costs, but because they can literally determine whether or not a life-saving medicine makes its way to patients. It’s impossible to know for sure, but many antibodies that could save lives seem not to make it to the clinic, simply because they are deemed “un-manufacturable” with existing tools.
We’re excited about our progress so far, but this is just the beginning.
Asimov Labs is already collecting multi-modal measurements on transcription and translation rates, protein folding, protein-protein interactions, and energy consumption for thousands of different DNA sequences, including promoters and terminators, in CHO cells. They are also designing new types of assays to study DNA context and plasmid architectures, including how the position in which DNA is inserted into the genome influences its ultimate expression. These datasets are then used to improve our engineering tools. Our modeling teams have also made a CHO simulator tool to predict how metabolite levels shift as cells make antibodies. (We’ll write about this in our next blog.)
In the next few years, we’d like to collect data and build models that can predict the optimal genetic design and bio-manufacturing conditions of a therapeutic molecule merely by looking at its sequence. Imagine if you could take an antibody sequence (say, for Humira®), type it into a computer, and then get an output that says, “Build this exact plasmid sequence, put it into a cell with this genome, and grow the cells under these conditions.”
Our CHO Edge tools are being used for antibodies first, but we plan to expand to materials, enzymes, and other molecules in the future.
If this work sounds exciting, come join us. Or, subscribe for updates.
Contributors: HaeWon Chung, Kevin Smith, Lila Wroblewska, Martín Cárcamo, Jamie Freeman, Scott Estes, Rachel Kelemen, Alec Nielsen & Ben Gordon. Words by Niko McCarty.


WHOO! Keep up the good work! Streamlining biologics manufacturing undoubtedly means (in time) a decrease in price for drugs that only have increases in demand. On behalf of humanity, and for all the people taking these drugs, THANK YOU!